Trading Places Review How Do Dna Polymerases Switch During Translesion Dna Synthesis?
Abstract
In response to UV irradiation, translesion DNA synthesis (TLS) utilizes specialized DNA polymerases to featherbed replication-blocking lesions. In a well-established polymerase switch model, Polη is thought to exist a preferred TLS polymerase to insert correct nucleotides across from the thymine dimer, and Rev1 plays a scaffold role through physical interaction with Polη and the Rev7 subunit of Polζ for continual DNA synthesis. Defective Polη causes a variant class of xeroderma pigmentosum (XPV), a disease with predisposition to sunlight-induced skin cancer. Previous studies revealed that expression of Rev1 alone is sufficient to confer enhanced UV harm tolerance in mammalian cells, which depends on its physical interaction with Polζ but is independent of Polη, a determination that appears to contradict current literature on the critical roles of Polη in TLS. To exam a hypothesis that the Rev1 catalytic activity is required to backup Polη in TLS, nosotros establish that the Rev1 polymerase-dead mutation is synergistic with either Polη mutation or the Polη-interaction mutation in response to UV-induced DNA impairment. On the other paw, functional complementation of polH cells by Polη relies on its physical interaction with Rev1. Hence, our studies reveal critical interactions between Rev1 and Polη in response to UV damage.
Introduction
Translesion DNA synthesis (TLS) is a means of Dna damage tolerance (DDT) that allows replication to bypass DNA damage via specialized DNA polymerases with or without associated increase in mutagenesisane,2. Mammalian TLS polymerases include Y-family Polη, Polκ, Polι and Rev1; they lack 3′–5′ proofreading exonuclease activity and replicate DNA in a distributive manneriii,iv. In addition, Polζ is a B-family TLS polymerase whose primary function is to extend Dna synthesis subsequently initial insertion past a Y-family polymerase opposite the damage site5,6. Although a Polζ2 complex containing a Rev3 catalytic subunit and a Rev7 regulatory subunit displays TLS polymerase activeness in vitrovii, an active Polζfour in vivo contains two Polδ subunitsviii,9.
The TLS response to UV irradiation has been extensively studied in mammalian cells. UV mainly causes 2 types of lesions: cyclobutane pyrimidine dimers (CPDs) and pyrimidine (six–4) pyrimidine photoproducts [(6–4)PPs]ten. Polη bypasses CPDs with high allegiance11,12, and defective Polη causes the variant form of the homo syndrome xeroderma pigmentosum (XPV) with increased risk of sunlight-induced skin cancer13,fourteen. Polη consists of a polymerase core region and a C-terminal domain (CTD), both of which are necessary for its biological functions15. The Polη-CTD contains a ubiquitin-binding motif (UBZ), a nuclear localization signal (NLS), 2 Rev1-interacting (RIR) motifs and 2 PCNA-interacting (PIP) motifs3,sixteen,17. Subsequently DNA impairment, PCNA is monoubiquitinated at the K164 rest18, which signals stalled replication forks19. While PIP and UBZ are to enhance interaction with monoubiquitinated PCNA, which is required for Polη to be recruited to the Deoxyribonucleic acid lesionsxvi, the role of RIR motifs in TLS remains controversial. It has been reported that Polη is required for the recruitment of Rev1 to the damage site through its RIR motifs, only ectopic expression of the RIR-defective Polη does not affect its ability to protect cells from UV-induced killing and mutagenesis17. Nevertheless, others reported that Rev1 and Polη are independently recruited to the damage site subsequently UV irradiation20,21. Furthermore, the PIP motif shares structural similarity with the defined RIR22 and indeed can interact with Rev123,24. Afterwards UV irradiation, Polη-CTD also promotes Rad18-mediated PCNA monoubiquitination that assists with the recruitment of error-prone TLS polymerases like Polι and Polκ25.
It has been well accepted that Rev1 functions as a scaffold for polymerase switch during TLS in response to UV irradiation26,27, in which its catalytic activeness is disposable, as the Rev1 polymerase-dead mutation does not confer increased sensitivity to UV-induced killing and mutagenesis28. Rev1 can exist recruited to the damage site through enhanced affinity for monoubiquitinated PCNA via its PCNA-binding BRCT domain29,xxx and ubiquitin-binding UBM motifs31. The Rev1-CTD contains 2 dissever domains to interact with Y-family polymerases including Polη, Polι and Polκ, and the Rev7 subunit of Polζ32,33,34,35. Nosotros recently attempted to address detailed scaffold roles of Rev1 in response to UV-induced DNA impairment and surprisingly found that UV harm tolerance conferred by ectopic expression of Rev1 is dependent on its interaction with Rev7 but independent of Polη interaction36. The current study further addressed genetic and concrete interactions between Rev1 and Polη, which immune the states to conclude that the Rev1 polymerase can play a backup role in the absenteeism of Polη and that Polη requires its RIR motifs to protect cells from UV-induced Deoxyribonucleic acid harm.
Results
Synergistic interaction between Rev1 polymerase and Polη-interaction mutations
We previously reported that UV damage tolerance mediated past PCNA-Ub fusion is dependent on Rev1 but independent of Polη37. Since Rev1 interacts with Polη and Rev7 through its Rev1-CTD that tin can be further divided into two subdomains34,35, we screened a large number of reported point mutations in this region34,35,38,39 and identified four mutations either specifically affecting Polη simply not Rev7 bounden (L1170A and V1188A), or disrupting Rev7 but not Polη binding (Y1242A and L1246A). By using an RPA nuclear focus formation analysis as an indication of TLS activity after UV irradiation37,40, it was constitute that Rev1-L1170A and Rev1-V1188A protected cells to a level comparable to that of Rev1, while CTD-Y1242A and CTD-L1246A lost Rev1 functions36 (Supplementary Fig. S1), which indicates that Ddt provided by Rev1 does not require its physical interaction with Polη.
The higher up observations are highly unexpected, equally it has been well established in both yeast and mammalian cells that Polη plays a critical function in TLS in response to UV irradiation, and that the Rev1-Polη interaction is critical during this procedurethree,41. Based on these observations, we wished to test a hypothesis that in the absence of Rev1-Polη interaction, Rev1 uses its ain catalytic activity to initiate TLS in response to UV-induced Deoxyribonucleic acid damage. To this end, we cloned Rev1-DE (Rev1-D568A, E569A, polymerase dead), Rev1-L1170A and the corresponding double mutant Rev1-DE-1170 into pEGFP-C1 equally GFP fusions. These plasmids were transfected into 293T cells, with vector pEGFP-C1 as a negative command and Rev1-GFP as a positive control, followed by two functional assays as previously described37. For the RPA nuclear focus germination assay, typical RPA2-positive- and negative-cells are illustrated in Fig. 1A. Compared with vector-transfected control, ectopic expression of Rev1 can significantly reduce the percent of cells with RPA2-positive foci later UV irradiation (Fig. 1B,C), and confer UV damage tolerance (Figs. 1D and S2A). Under the above experimental weather condition, expression of either Rev1-DE or Rev1-1170 is sufficient to bring the percent of RPA2-positive cells to the wild-type level (Fig. 1B,C) and confer near wild-type level UV tolerance (Fig. 1D). In sharp contrast, ectopic expression of the Rev1-DE-1170 double mutant did not confer UV tolerance in either assay in comparing to the control transfected cells (Fig. 1). We ruled out the possibility that the lack of DNA-damage tolerance function of Rev1-DE-1170 was due to contradistinct factor expression or protein stability (Fig. 2A). The in a higher place observations collectively allow us to conclude that the Rev1 polymerase and Polη-interaction mutations are synergistic in response to UV-induced Deoxyribonucleic acid damage.
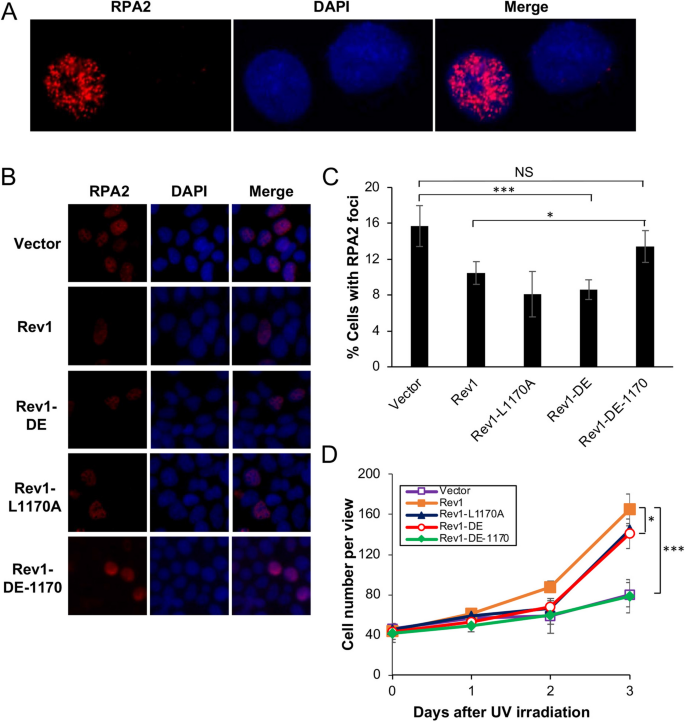
Effects of Rev1 and its mutant derivatives on cellular tolerance to UV irradiation in 293T cells. (A, B) Representative images of an RPA nuclear focus formation analysis. 293T cells were transfected with plasmids expressing GFP-Rev1 or its mutations. 48 h afterward, these cells were irradiated past 8 J/m2 UV and incubated for 6 h before staining with DAPI or an antibiotic against RPA2. (A) represents typical RPA2-positive (left) and RPA2-negative (right) cells. (C) Quantitative analysis of data from (B). (D) Effects of Rev1 and its mutations on 293T cell growth in response to UV irradiation. Cells were transfected with plasmids expressing wild-type or indicated Rev1 bespeak mutations for 2 days before 30 J/1000ii UV irradiation. (C, D) Data are ways of three contained experiments ± SEM. *, P < 0.05; ***, P < 0.001; NS, not significant past 2-sided Student'south t test.
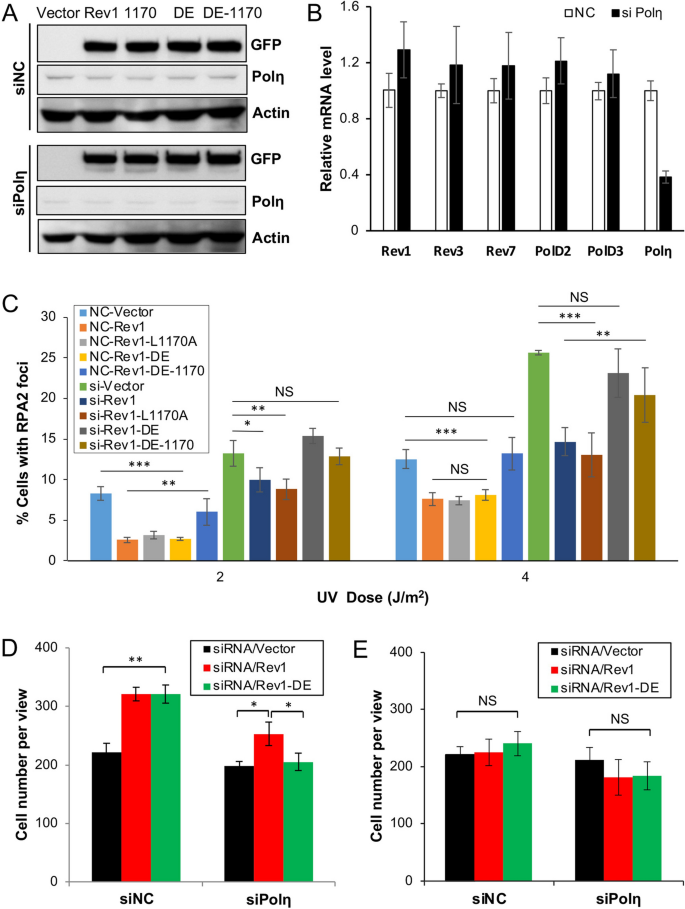
Rev1 and Polη play alternative roles at the insertion step of TLS. (A) Western blot assay of GFP-mRev1 and its mutant transfectants in 293T cells. Cells were transfected with siPolη or non-specific siRNA (siNC). 24 h afterwards, these cells were transfected with plasmids expressing GFP-mRev1 or its mutant proteins. Later 48 h, the transfected cells were harvested, lysed and subjected to western blotting. The two sets of gels were from the same experiment and treated under identical conditions. (B) Efficacy of siRNA depletion against Polη in 293T cells as measured by qRT-PCR assay. (C) Effects of GFP-mRev1 or its mutant expression on UV-induced nuclear RPA2 focus formation in 293T cells with siPolη (si) or non-specific siRNA (NC) treatment. Cells were transfected with siRNA molecules in combination with GFP-mRev1 or its mutants followed past UV irradiation. Immunofluorescence assay was performed 6 h after UV irradiation. (D, E) Furnishings of Polη depletion and ectopic expression of GFP-mRev1 or GFP-mRev1-DE on 293T cell growth with (D) or without (E) UV irradiation. Cells were transfected with siPolη. 24 h afterward, these cells were transfected with GFP-mRev1 or its mutant plasmids, incubated for 2 days and irradiated by 20 J/thousand2 UV. Later on 72 h of incubation, the number of feasible cells were counted. Data shown in (A, C, D, E) are means of at least iii independent experiments ± SEM. *, P < 0.05; **, P < 0.01; ***, P < 0.001; NS, not meaning by two-sided Student'south t examination.
Rev1 and Polη play alternative roles at the insertion step of TLS in bypassing UV-induced lesions
The Rev1-L1170A mutation likely affects interaction with all three Y-family unit polymerases, namely Polη, Polι and Polκ32,33. Since Polη plays a critical part in cellular response to UV irradiation, nosotros hypothesized that the Rev1 catalytic activity is to back up Polη in the Rev1-L1170A background. To test this hypothesis, we asked whether compromised Polη could supervene upon the Rev1-L1170A mutation. To this end, we depleted the endogenous Polη past siRNA to approximately sixteen% of the wild-type level (Fig. S3, likewise see Fig. 4B) while expressing GFP-REV1 and its various mutations. GFP-REV1 and its mutants expressed equally well in siPolη cells in comparing to not-specific siNC cells (Figs. 2A and S3), while siPolη specifically reduced the transcript level of the POLH factor, simply not other relevant genes encoding Rev1 and Polζ subunits (Fig. 2B). After depletion of Polη from 293T cells, the percentage of RPA2-positive cells about doubled that of the control group, while ectopic expression of Rev1 or Rev1-1170 caused a subtract in RPA2-positive cells to the same extent. Interestingly, expression of Rev1-DE alone provided UV resistance to siNC cells, only failed to protect Polη-depleted cells, in which the per centum of RPA2-positive cells was similar to that of Rev1-DE-1170-transfected cells (Fig. 2C). The in a higher place observations point that the Polη depletion is epistatic to Rev1-1170A and condiment to Rev1-DE. The additive effect between Polη depletion and the Rev1-DE mutation was also seen in a 20 J/grand2 UV-induced cell survival analysis, in which Rev1-DE-transfected cells behave like Rev1 afterward siNC treatment, but like empty vector afterwards siPolη handling (Fig. 2D,Due east). Based on the to a higher place observations, we infer that Rev1 and Polη play alternative roles at the insertion pace of TLS upon UV irradiation.
UV damage tolerance conferred by Polη is partially dependent on its interaction with Rev1
Our observations that the Rev1 interaction with Polη is dispensable appear to contradict a notion of functional importance of concrete interaction between Rev1 and other Y-family unit polymerases during TLS3,41. One possibility is that our written report was under the Rev1 ectopic expression condition, in which excessive Rev1 is sufficient to provide fill-in catalytic activity during TLS. It has been previously reported that Polη interacts with Rev1 through residues 369–49120 and 509–55733, designated equally RIR1 and RIR2, respectively (Fig. 3A), and that an FF motif is critical for this interaction22. We made RIR1 (Polη-FF483,484AA), RIR2 (Polη-FF531,532AA) and the corresponding double mutation RIRD, and examined their furnishings on Polη functions. Under our experimental conditions, ectopic expression of GFP-POLH and its mutant forms resulted in approximately sixfold more than GFP-Polη over endogenous Polη, as judged by western blot analysis (Fig. 3B). GFP-Polη transfection reduced UV-induced RPA2-positive cells (Fig. 3C,D) and protected cells from killing by UV (Figs. 3E and S2B). We then assessed whether the UV damage tolerance conferred by Polη was dependent on its interaction with Rev1.
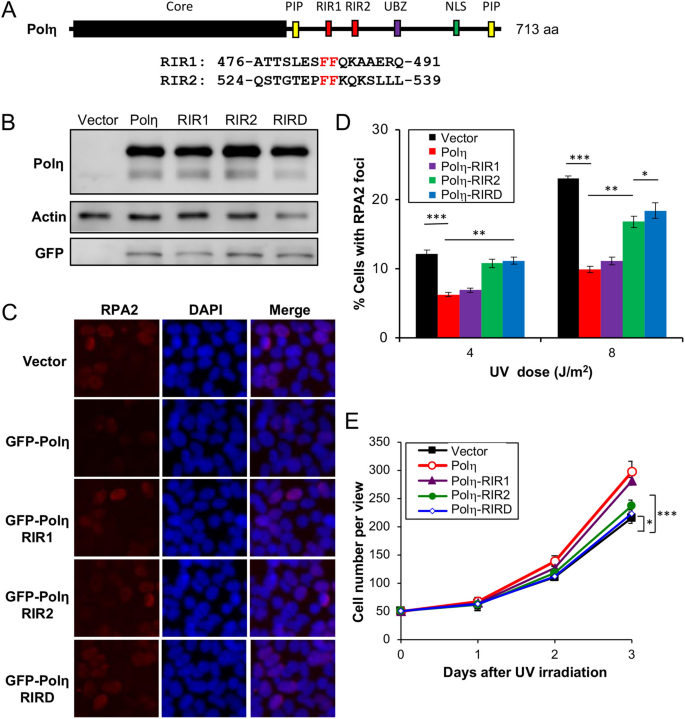
Effects of Polη and its mutation derivatives on cellular tolerance to UV irradiation in T-King-293 cells. (A) Illustration of the Polη structure. Core, the Y-family polymerase catalytic domain; PIP, PCNA-interaction peptide; RIR, Rev1-interaction region; UBZ, Ub zinc-finger; NLS, nuclear localization point. The RIR1 and RIR2 sequences are aligned with the consensus FF residues in red. (B) Western blot assay of GFP-Polη transfectants. (C) Representative images of an RPA nuclear focus germination assay after eight J/m2 UV irradiation. (D) Quantitative analysis of the RPA nuclear focus formation assay after 6 h of incubation following UV irradiation. (E) Effects of Polη and its mutations on cell growth in response to UV irradiation. Cells were transfected with plasmids expressing wild-type or POLH point mutations and incubated for 2 days before 30 J/mtwo UV irradiation and counting viable cells over fourth dimension. Results in (D, E) are means of iii independent experiments ± SEM. *, P < 0.05; **, P < 0.01; ***, P < 0.001 by two-sided Educatee's t test.
The Polη-RIR1 mutation seemed to have a moderate effect on the resistance provided by Polη. In contrast, the Polη-RIR2 mutation had a dramatic consequence on the Polη part in both RPA2 foci (Fig. 3C,D) and jail cell survival (Fig. 3E) assays. When T-REx-293 cells were transfected with Polη in which both RIR motifs were mutated, the UV-induced RPA2-positive cells farther increased over the level in Polη-RIR2 transfected cells (Fig. 3C,D), indicating that Polη-RIR1 contributes moderately to Rev1 bounden. The above observations collectively let us to conclude that the Polη-Rev1 interaction is mainly through the Polη-RIR2 motif, and that UV impairment tolerance conferred by Polη overexpression appears to be partially dependent on its interaction with Rev1.
POLH lacking cells were sensitive to UV-induced Dna impairment
Loss of Polη activity is responsible for the XPV cells13,14. To further investigate the role of Rev1 and Polη in TLS, we established POLH-inactivated cell lines past knocking out the XPV/POLH factor from 293T cells using a CRISPR/Cas9 method42,43,44. 1 of the cell lines, POLH-1, contains a homozygous 2-bp deletion at the second exon, causing a frameshift mutation (Fig. 4A). A western blot analysis compared endogenous Polη levels in 293T cells, siPolη-treated cells and the isogenic POLH-one cells. While siPolη handling reduced cellular Polη past 84%, Polη is undetectable in the POLH-1 cells (Fig. 4B). Compared with the parental 293T cells, POLH-1 cells displayed a relatively normal proliferation rate in the absence of UV irradiation; however, upon 5 J/thousand2 UV irradiation, the POLH-1 cells stopped proliferation over 3 days, whereas the proliferation of 293T cells was only moderately affected (Fig. 4C). 293T cells displayed a feature increase in RPA2-positive cells with increasing doses of UV irradiation, reaching approximately 20% at 8 J/m2. In contrast, POLH-one cells dramatically increased RPA2-positive cells to over 40% upon ii J/gtwo UV irradiation and did non further increase with increasing doses of UV (Fig. 4D), probably because most cells were dead. two J/yard2 UV irradiation did not significantly induce RPA2 focus formation in 293T cells over time, but drastically induced RPA2 focus germination in POLH-1 cells within 2 h, and the percentage of RPA2-positive cells gradually increased over time (Fig. 4E). The above results confirmed the successful establishment of a polH null cell line and demonstrated that POLH-1 cells sustain UV-induced ssDNA as a hallmark of defective TLS.
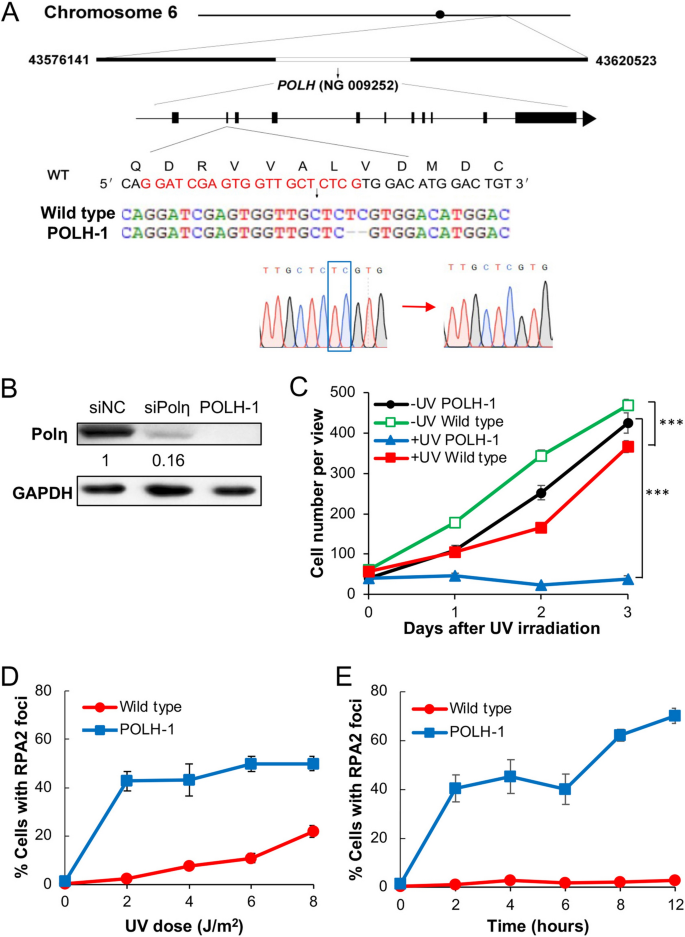
Creation and characterization of an POLH-1 prison cell line. (A) The POLH cistron location, genomic construction and mutation in the POLH-1 cell line. Nucleotide and encoded amino acid sequences around the guide RNA target (in red) are shown. DNA sequence confirmation of the 2-nt deletion (blue box) in the POLH-ane cells is as well illustrated. (B) Western blot analysis of Polη in 293T, siPolη-treated and POLH-1 cells. For siPolη depletion, 293T cells were transfected with siPolη molecules and harvested 48 h after treatment. The number indicates the band intensity relative to non-specific siRNA (siNC) treated cells. (C) Relative cell growth with or without UV irradiation. 293T and POLH-1 cells were cultured for 2 days followed by 5 J/gtwo UV irradiation and counting feasible cells over time. (D) RPA2 focus formation in 293T and POLH-ane cells 4 h after exposure with unlike UV doses. (E) RPA2 focus formation in 293T and POLH-1 cells after ii J/m2 UV irradiation over fourth dimension. (C-East) Data are means of iii independent experiments ± SEM. ***, P < 0.001 by two-sided Educatee'south t test.
Effects of Rev1 and its mutant derivatives on tolerance to UV irradiation in POLH-i cells
Previously, we hypothesized that the catalytic action of Rev1 plays a role in nucleotide insertion during TLS either when Rev1 cannot interact with Polη or when the endogenous Polη is reduced. Withal, the above results are discipline to a different estimation, every bit they were not obtained from a strict genetic system. With the creation of POLH-1 cells, we were able to critically test our original hypothesis in a clean genetic groundwork. Indeed, in comparing to vector-transfected cells, ectopic expression of REV1 in POLH-1 (Fig. 5A) could reduce UV-induced RPA2-positive cells (Fig. 5B,C). Nether the above experimental conditions, expression of REV1-L1170A could rescue POLH-1 cells to the wild-type REV1 level, whereas expression of REV1-DE or the double mutation was no longer able to protect POLH-ane cells (Fig. 5B,C). Similarly, expression of REV1 or REV1-L1170A protected POLH-ane cells from killing by UV irradiation to the same level, while expression of REV1-DE or REV1-DE-1170 had no protective effect (Fig. 5D). These results, together with previous observations36, conspicuously show that when REV1 is overexpressed, its Rev7 interaction is absolutely required for cellular tolerance against UV damage, while either its catalytic action or Polη, merely not both, is dispensable.
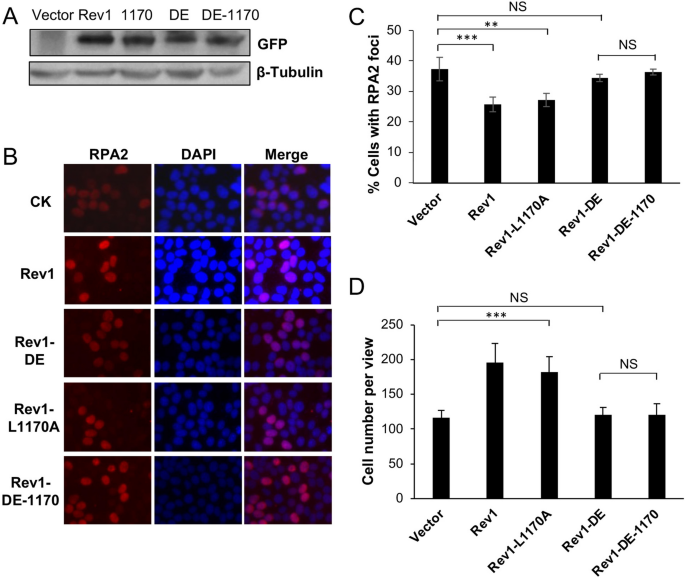
Effects of mRev1 and its mutant derivatives on tolerance to UV irradiation in POLH-1 cells. (A) Western blot analysis of transfected GFP-Rev1 and its mutations in POLH-1 cells. (B) Representative images of an RPA nuclear focus assay. POLH-1 cells were transfected with plasmids expressing GFP-Rev1 or its mutations for two days followed by 2 J/m2 UV irradiation and incubation for 6 h before staining with DAPI or an antibody against RPA2. (C) Quantitative analysis of the RPA2 nuclear focus germination. (D) Furnishings of Rev1 and its mutations on POLH-one cell growth in response to UV irradiation. Cells were transfected with plasmids expressing GFP-Rev1 or its betoken mutations for two days before 2 J/m2 UV irradiation, then these cells were cultured for 48 h, followed by counting viable cells. (C, D) Results are means of three independent experiments ± SEM. **, P < 0.01; ***, P < 0.001, NS, non significant by 2-sided Student's t test.
Effects of Polη and its mutant derivatives on tolerance to UV irradiation in POLH-ane cells
Concrete interaction betwixt Polη and Rev1 may facilitate TLS; however, it is unclear whether Polη-mediated DDT can bypass the requirement for Rev1. To examination whether RIR mutations affect the analogousness of Polη for Rev1, nosotros co-transfected POLH-1 cells with GFP-Rev1-CTD and Polη-RIR mutant derivatives followed by co-IP against GFP and western blot analysis against Polη. Figure 6A shows that, compared to wild-type Polη, the Polη-RIR2 mutation had a stronger effect on binding to GFP-Rev1-CTD than the Polη-RIR1 mutation, and the Polη-RIRD double mutation further reduced its affinity for GFP-Rev1-CTD. Hence, RIR2 appears to play a major role in the Polη-Rev1 interaction. The remaining coimmunoprecipitated Polη may come from indirect interactions, as both Rev1 and Polη interact with PCNA and Ubiii,xvi,29,31, or from other putative RIR motifs found in Polη22.
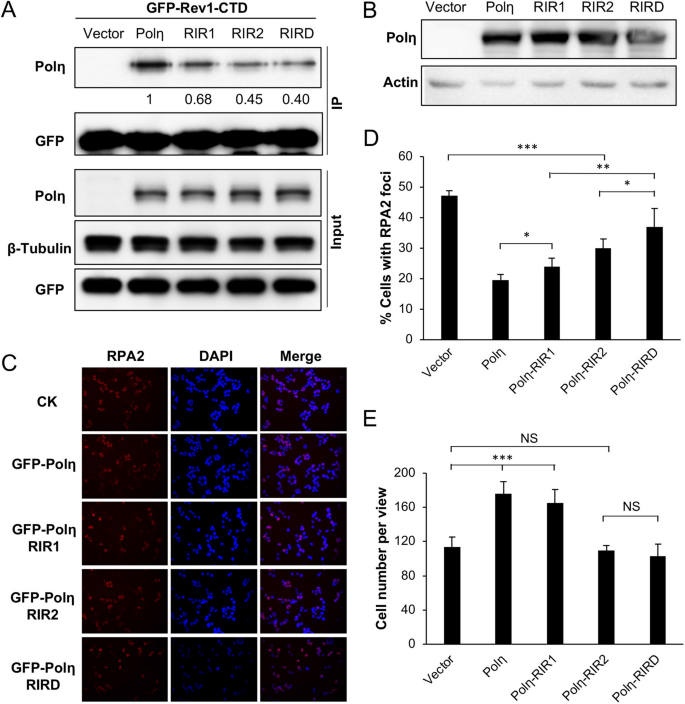
Effects of Polη and its mutant derivatives on UV damage response in POLH-i cells. (A) Co-IP assays to appraise the interaction between Rev1 and mutated Polη in POLH-1 cells. (B) Western blot analysis of ectopic expression of POLH and its mutations in POLH-1 cells. (C, D) Effects of Polη and its mutations on preventing UV-induced RPA2 nuclear focus germination. POLH-1 cells were transfected with the indicated plasmids and and then incubated for 2 days earlier 4 J/chiliad2 UV irradiation, followed by continued civilisation for 6 h and staining with DAPI or an antibiotic against RPA2. (C) Representative images of the RPA nuclear focus assay. (D) Quantitative analysis. (E) Effects of Polη and its mutations on POLH-one cell growth in response to UV irradiation. Cells were transfected with plasmids expressing Polη or its point mutations earlier 2 J/one thousandii UV irradiation, and so these cells were cultured for 48 h, followed by counting viable cells. (D, East) Results are means of three contained experiments ± SEM. *, P < 0.05; ** P < 0.01; ***, P < 0.001, NS, non pregnant by two-sided Student's t examination.
We transfected plasmids producing GFP-Polη and its RIR mutant forms in POLH-1 cells and monitored their response to UV irradiation. Mutant forms of GFP-Polη did not bear upon their expression and protein stability in POLH-i cells under our experimental weather condition (Fig. 6B). Expression of POLH in POLH-one reduced UV-induced RPA2-positive cells to a great extent, and expression of POLH-RIR1 and POLH-RIR2 mutations partially restored the Polη part in POLH-one cells, while expression of the POLH-RIRD double mutant grade of Polη further reduced its rescue ability (Fig. 6C,D). Hence, Polη-RIR1 and Polη-RIR2 mutations appear to be condiment in affecting Rev1 interaction, which plays a critical office in TLS. In a cell survival assay, expression of POLH restored POLH-1 cell tolerance to 2 J/kii UV and expression of POLH-RIR1 had a like effect. In contrast, expression of either POLH-RIR2 or POLH-RIRD failed to rescue POLH-1 cells from killing by UV irradiation (Fig. 6E), indicating that the RIR2 motif plays a critical office in Rev1 interaction and is absolutely required during TLS in response to UV under our experimental conditions. These observations collectively permit us to conclude that interaction with Rev1 is critical for Polη to role in response to UV irradiation.
Discussion
In mammalian cells, PCNA plays crucial roles in Dna replication and repair45. PCNA interacts and travels with all three replicative polymerases during chromosomal DNA replication. When DNA damage stalls the replication fork, PCNA tin exist ubiquitinated at its K164 residue past Rad6-Rad18, switching to a Dichloro-diphenyl-trichloroethane mode18. Monoubiquitinated PCNA enhances affinity for Y-family polymerasessixteen including Polη, Polι, Polκ and Rev1, all of which contain PCNA- and Ub-binding domains46. In response to UV irradiation, both Polη and Rev1 are colocalized to the damage sites in the form of nuclear focixx. Although subject to debate, our own observations21,36 favor a previous study20 that they are recruited to the damage site independently from each other, which raises a critical effect: what is the office of Rev1-Polη interaction during TLS? We previously reported that UV damage tolerance conferred past ectopic expression of PCNA-Ub fusion37 and Rev136 depends on Rev1 and its physical interaction with Polζ, respectively, simply is independent of Polη. Hither nosotros testify that ectopic expression of Polη tin can confer boosted UV harm tolerance, which requires its RIR domains. Furthermore, expression of Polη tin rescue the increased UV sensitivity in Polη-defective cells that mimic the XPV syndrome. This rescue relies on Polη'southward concrete interaction with Rev1 through RIR motifs. Our observations differ from a previous report17 that the Polη-RIR mutations do not affect Polη rescue of XPV cells from killing past UV irradiation, but are consistent with a report47 that expression of a polymerase-dead Polη moderately rescues UV sensitivity of Polη-aught mouse cells, which depends on its interaction with Rev1. Although both RIR motifs accept been reported to mediate interaction with Rev1twenty,33, we found that RIR2 plays a disquisitional role while RIR1 may play a fill-in role, although RIR1 is critical for the interaction with PolD248. In the absence of RIR motifs, Polη nevertheless retains certain physical interaction with Rev1, probably through cryptical RIR motifs institute in Polη22, although it is insufficient to support the Polη TLS activity. We propose a matchmaker mechanism in which only when cells sense the presence of both Polη and Rev1 at the aforementioned impairment site through their concrete interaction volition insertion by Polη and extension by Rev1-mediated Polζ take place to consummate the ii-pace TLS (Fig. 7).
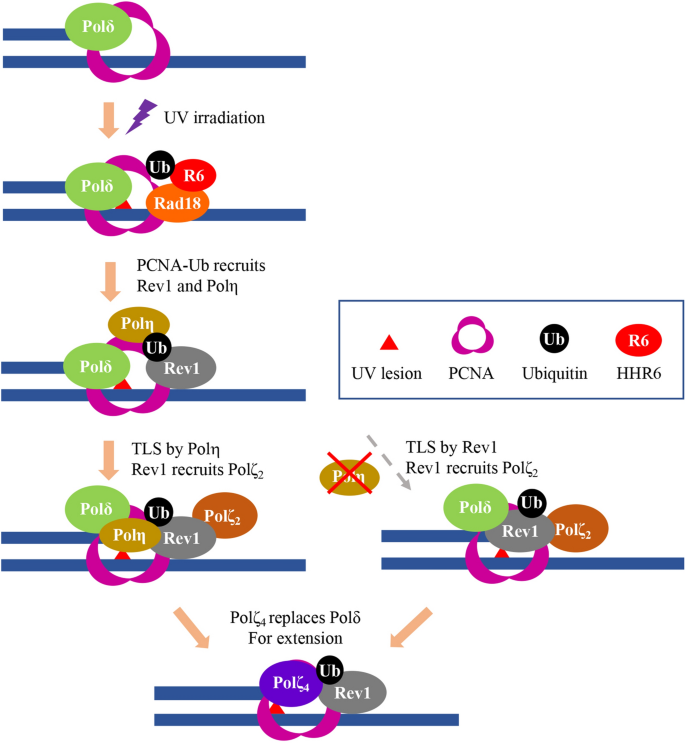
Proposed working model for TLS in response to UV irradiation in mammalian cells. UV irradiation induced DNA impairment blocks replicative polymerase like Polδ. The ssDNA along with stalled replication fork recruits the Rad6-Rad18 complex to monoubiquitinate PCNA-K164, which in plow recruits both Polη and Rev1. A default pathway is for Polη to insert nucleotides opposite the lesion, and for Rev1 to recruit Rev7-Rev3 (Polζtwo) to the damage site to form Polζfour for extension, in which the Polη-Rev1 interaction plays a crucial office. In the absence of Polη, Rev1 can play dual roles in both insertion and Polζ2 recruitment, in which its catalytic action is required.
It has been well accepted that in response to UV irradiation, Polη plays a critical catalytic role while Rev1 only plays a scaffold office; the catalytic activity of Rev1 is involved in bypassing abasic sites49 and lesions induced by 4-nitroquinoline-1-oxide50,51,52, but is dispensable for lesions induced past UV28. However, some studies have indicated direct or indirect roles of Rev1 in bypassing CPD and (6–4)PP53,54,55. Since UV damage tolerance conferred by Rev1 is independent of its physical interaction with Polη36, we critically tested a hypothesis that the Rev1's catalytic activity is responsible for the observed Ddt. Firstly, nosotros found a strong synthetic effect between the catalytic and Polη-binding mutations in Rev1. While either the Rev1-DE or Rev1-L1170A mutation tin provide Ddt to nigh the wild-type Rev1 level, the double mutation behaves like a zero mutation. Secondly, nosotros ruled out a possibility that other TLS polymerases are responsible for the effect past experimental depletion of Polη, and found that one time Polη is depleted, the Rev1-L1170 mutation still behaves like wild-type Rev1, while the Rev1-DE mutation is no longer able to provide Dichloro-diphenyl-trichloroethane. Finally, using a newly created POLH-1 cell line, we showed that the polh is additive with the Rev1-DE mutation just epistatic to the Rev1-L1170A mutation. These observations collectively support a notion that Polη plays a frontline role in TLS insertion across from UV-induced lesions and that Rev1 is required for its non-catalytic office, probably through recruiting Polζii. However, in the absence of Polη, Rev1 can insert nucleotide(s) across from UV-induced lesions equally well every bit recruit Polζ2 to the impairment site (Fig. 7). A major critique for the above model is that it is based on ectopic expression of Rev1, which may not occur in untransfected cells. Interestingly, a synergistic interaction between Rev1-DE and Polη in response to UV irradiation is recently reported in budding yeast56, which lends stiff support to the working model. Rev1 may likewise facilitate the assembly of Polζiv by physical interaction with PolD357, which is likely an active form for TLS extension58 (Fig. 7). Rev1 is a template-dependent dCMP transferase51,59. Since UV-induced lesions are nigh exclusively on pyrimidines60, the dCMP insertion by Rev1 is expected to cause transversion mutations. Fortunately Rev1 has very limited catalytic activity toward major UV-induced lesions52, and it must exist kept at bay until needed.
Methods
Plasmids and plasmid construction
The open up reading frames of POLH and mRev1 were cloned in pEGFP-C1 (BD Biosciences Clontech). Polη point mutants were created with the following primers: RIR1-F: 5′-ACCACGTCTGGAATCAGCCCAAAGCTGCAGAAAGG-three′; RIR1-R: v′-CCTTTCTGCAGCTTTGGGGGGCTGATTCCAGACGTGGT-3′; RIR2-F: v′-AGTACAGGAACTGAGCCCGCTAAGCAAAAGTCTGCTT-3′; RIR2-R: 5′-AAGCAGACTTTTCT TG TAGCGGGCTCAGTTCCTGTACT-three′. Rev1 point mutants were created with the following primers: Rev1-L1170A-F: five′-AGTGATGTGAAGACCTTGGCCAAAGAGTGGATCACTACT-3′; Rev1-L1170A-R: 5′-AGTAGTGATCCACTCTTTGGCCAAGGTCTTCACATCACT-3′; Rev1-DE-F: 5′-ATCGAGGCTGTCAGCTGCGCTGCAGCACTGATTGACGTCACG-3′; Rev1-DE-R: v′-CGTGACGTCAATCAGTGCTGCAGCGCAGCTGACAGCCTCGAT-three′.
For the co-IP assay, DNA sequences corresponding to Rev1-CTD (residues 1150–1249) were PCR-amplified and cloned into pEGFP-C1 to produce EGFP-fused proteins. POLH and its mutant coding sequences were cloned into vector pcDNA4/TO (Invitrogen).
Prison cell civilisation and reagents
Human T-REx293 and 293T cells were purchased from Invrogen, and POLH-one cells were created from 293T cells in this study. Cells were cultured in a DMEM medium supplemented with 10% fetal bovine serum at 37 °C in the presence of 5% COtwo. For transient transfection experiments, T-REx293 and 293T cells were transfected with indicated plasmids by using PEI (Polyethylenimine, Linear, MW 25,000, Polysciences) following the manufacturer's protocols. In order to enrich transfected cells over 50%, G418 was added to a final concentration of 200 µg/mL 24 h after transfection. POLH-1 cells were transfected with indicated plasmids by using Lipofectamine 2000 (Invitrogen) to attain xl–50% transfection efficiency without subsequent antibiotic selection.
Generation of polh cell lines from 293T cells
CRISPR/Cas9-mediated factor targeting was performed by using a Genloci CRISPR/Cas9 kit with EGFP + Puror (GP0129, Genloci) as described (Protocol No. PT161117-1). Briefly, the POLH-targeting double-strand oligonucleotide, fabricated by annealing Polη-F: 5′-caccGGATCGAGTGGTTGCTCTCG-3′ and Polη-R: 5′- aaacCGAGAGCAACCACTCGATCC-3′, was cloned into plasmid pGK1.2. The resulting plasmid was used to transfect 293T cells, and the transfectants were cultured in a DMEM medium supplemented with ten% fetal bovine serum. Puromycin (Sigma) was used to a final concentration of 1 µg/mL to select transfectants over xiv days, and puromycin-resistant clones were transferred to a 96-well plate for expansion and screening of POLH knockouts. The targeted clones were confirmed by genomic PCR with primers Polη Primer-F: 5′-CCATGCTCCCATGCTCATGGTAACTC-3′ and Polη Primer-R: 5′-CCTGCCACAGTGCCACTGTGTTACC-3′, and the PCR products were sent for sequencing.
RNA interference
The depletion of endogenous Polη in 293T cells was performed equally previously described37. The POLH factor-specific target sequence (siPolη) 5′-CTGGTTGTGAGCATTCGTGTA-3′ and the scrambled siRNA (siNC) were purchased from Shanghai GenePharm. The suppression efficacy was assessed past quantitative RT-PCR (qRT-PCR) and/or western blotting 48 h after siRNA transfection. Primers used for qRT-PCR include RT-Polη-F: 5′-GCAGCCATAGAGAGGGAGAC-3′, RT-Polη-R: 5′-CTCCTTAATGTCACGCACGAT-three′, hRev1-F: 5′-ACCGAAGAGGAGCACAAAGA-iii′, hRev1-R: 5′-CCATTCCATTTCCCTGAAGA-3′, hRev3-F: v′-AGTAAATGTCGGAGCCAAC-3′, hRev3-R: 5′-CTGGGCAGTTCAGAGAAACA-3′, hRev7-F: v′-TGGCTGTGCATCTCATCCTCT-3′, hRev7-R: 5′-GCGGTGCTCTTTATCCAAAATCA-iii′, hPolD2-F: five′-CCATCAGCCAACAATGCCAC-3′, hPolD2-R: 5′-CTAGCCGGAAGGGTTGTGA-3′, hPolD3-F: 5′-GAGTTCGTCACGGACCAAAAC-iii′, and hPolD3-R: 5′-GCCAGACACCAAGTAGGTAAC-3′.
Cell survival assay
The 293T or POLH-ane cells were cultured in 6-cm civilisation dishes, and and then transfected with plasmids carrying the gene of interest. After incubation for 48 h, the cells were irradiated with UV at the given doses, cultured for up to 3 days and and then stock-still with 4% formaldehyde. The fixed cultures were stained with DAPI and photographs were taken from random fields in dish for cell counting. Cells with round and intact nuclei were counted as feasible cells, and images were acquired using the CCD RoHs (Q26053) as previously described37. At to the lowest degree 2000 cells were counted for each treatment.
RPA nuclear focus formation assay
Cultured cells were seeded on poly-lysine-coated comprehend slips, rinsed once with ice-cold PBS (2.25 one thousand Na2HPOiv, 8 g NaCl, 0.2 g KHtwoPO4, two g KCl dissolved in i L ddHiiO), treated with 0.four% NP-xl in PBS for xx min on water ice, and so fixed with 4% paraformaldehyde for 15 min. The fixed cells were rinsed 3 times with PBS, treated with methanol for five min, so rinsed 4 times with PBST for 5 min each time. Later incubation with five% FBS in PBST for 45 min, cells were incubated with mouse anti-Replication protein A2 (RPA2) antibody (Abeam, Ab2175, 1:grand) overnight. The cells were washed four times with PBST, incubated with Alexa Flour 546 goat anti-mouse secondary antibiotic (Invitrogen, A11030, 1:1000) and 1.5 µg/mL four′,6-diamidino-2-phenylindole (DAPI) at room temperature for 1 h, and finally washed iv times with PBST once again. For quantitative analysis of UV-induced RPA2 focus formation, the 293T or POLH-ane cells transfected with corresponding plasmids were treated with UV. Images were taken with the same exposure time. Microscopy was performed with an inverted Olympus 10*22 microscope equipped with a 40 × immersion lens, and images were acquired using the CCD RoHs (Q26053) as previously described37. At least 1000 cells were counted for each treatment.
Co-immunoprecipitation (Co-IP) and western blotting
To measure out the expression levels of GFP-Rev1 and Polη or their mutant derivatives, 293T cells were transfected with the plasmids. ii days later, jail cell lysates were harvested, boiled before SDS-Page, and detected by indicated antibodies against Polη (Abcam, ab17725) and the GFP Tag (Abmart, 7G9). The following reference protein antibodies were from Lifetech: GAPDH (GA331), β-Actin (GA321) and β-tubulin (GA311).
For the co-IP assay, the POLH-i cells transfected with GFP-Rev1-CTD and Polη RIR mutants were harvested and immunoprecipitated with GFP-Trap A (ChromoTek, gta-xx) overnight. The input and the immunoprecipitated proteins were separated past SDS-PAGE and proteins of involvement were detected past indicated antibodies confronting Polη, β-tubulin and GFP.
References
-
Shachar, S. et al. Two-polymerase mechanisms dictate fault-complimentary and error-prone translesion DNA synthesis in mammals. EMBO J. 28, 383–393. https://doi.org/x.1038/emboj.2008.281 (2009).
-
Zhang, West., Qin, Z., Zhang, Ten. & Xiao, W. Roles of sequential ubiquitination of PCNA in Deoxyribonucleic acid-damage tolerance. FEBS Lett. 585, 2786–2794. https://doi.org/10.1016/j.febslet.2011.04.044 (2011).
-
Yang, W. & Woodgate, R. What a difference a decade makes: insights into translesion Dna synthesis. Proc. Natl. Acad. Sci. U.s.A. 104, 15591–15598. https://doi.org/10.1073/pnas.0704219104 (2007).
-
Andersen, P. L., Xu, F. & Xiao, Due west. Eukaryotic Dna damage tolerance and translesion synthesis through covalent modifications of PCNA. Cell Res. 18, 162–173 (2008).
-
Johnson, R. E., Washington, K. T., Haracska, 50., Prakash, S. & Prakash, L. Eukaryotic polymerases iota and zeta act sequentially to bypass Dna lesions. Nature 406, 1015–1019. https://doi.org/10.1038/35023030 (2000).
-
Prakash, S. & Prakash, L. Translesion DNA synthesis in eukaryotes: a one- or two-polymerase thing. Genes Dev. 16, 1872–1883. https://doi.org/10.1101/gad.1009802 (2002).
-
Nelson, J. R., Lawrence, C. W. & Hinkle, D. C. Thymine-thymine dimer bypass by yeast DNA polymerase zeta. Science 272, 1646–1649. https://doi.org/10.1126/scientific discipline.272.5268.1646 (1996).
-
Johnson, R. Eastward., Prakash, L. & Prakash, S. Pol31 and Pol32 subunits of yeast DNA polymerase delta are too essential subunits of DNA polymerase zeta. Proc. Natl. Acad. Sci. U.S.A. 109, 12455–12460. https://doi.org/10.1073/pnas.1206052109 (2012).
-
Makarova, A. V., Stodola, J. L. & Burgers, P. M. A four-subunit DNA polymerase zeta complex containing Pol delta accessory subunits is essential for PCNA-mediated mutagenesis. Nucleic Acids Res. 40, 11618–11626. https://doi.org/ten.1093/nar/gks948 (2012).
-
Sinha, R. P. & Hader, D. P. Physiological aspects of UV-excitation of DNA. Acme. Curr. Chem. 356, 203–248. https://doi.org/x.1007/128_2014_531 (2015).
-
Johnson, R. E., Prakash, S. & Prakash, L. Efficient bypass of a thymine-thymine dimer by yeast DNA polymerase, Poleta. Science 283, 1001–1004. https://doi.org/ten.1126/scientific discipline.283.5404.1001 (1999).
-
Biertumpfel, C. et al. Construction and mechanism of human Deoxyribonucleic acid polymerase eta. Nature 465, 1044–1048. https://doi.org/10.1038/nature09196 (2010).
-
Johnson, R. E., Kondratick, C. M., Prakash, S. & Prakash, L. hRAD30 mutations in the variant course of xeroderma pigmentosum. Science 285, 263–265. https://doi.org/10.1126/science.285.5425.263 (1999).
-
Masutani, C. et al. The XPV (xeroderma pigmentosum variant) factor encodes human Dna polymerase eta. Nature 399, 700–704. https://doi.org/10.1038/21447 (1999).
-
Prakash, S., Johnson, R. East. & Prakash, L. Eukaryotic translesion synthesis DNA polymerases: specificity of structure and function. Annu. Rev. Biochem. 74, 317–353. https://doi.org/10.1146/annurev.biochem.74.082803.133250 (2005).
-
Bienko, M. et al. Ubiquitin-binding domains in Y-family polymerases regulate translesion synthesis. Scientific discipline 310, 1821–1824. https://doi.org/ten.1126/science.1120615 (2005).
-
Akagi, J. et al. Interaction with DNA polymerase eta is required for nuclear accumulation of REV1 and suppression of spontaneous mutations in human cells. DNA Repair (Amst.) viii, 585–599. https://doi.org/10.1016/j.dnarep.2008.12.006 (2009).
-
Hoege, C., Pfander, B., Moldovan, G. Fifty., Pyrowolakis, K. & Jentsch, S. RAD6-dependent Dna repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419, 135–141. https://doi.org/10.1038/nature00991 (2002).
-
Pastushok, 50. & Xiao, W. Dna postreplication repair modulated past ubiquitination and sumoylation. Adv. Protein Chem. 69, 279–306 (2004).
-
Tissier, A. et al. Co-localization in replication foci and interaction of human Y-family members, Dna polymerase politico eta and REVl poly peptide. Dna Repair (Amst.) 3, 1503–1514. https://doi.org/10.1016/j.dnarep.2004.06.015 (2004).
-
Andersen, P. L., Xu, F., Ziola, B., McGregor, West. G. & Xiao, W. Sequential associates of translesion DNA polymerases at UV-induced DNA damage sites. Mol. Biol Jail cell 22, 2373–2383. https://doi.org/x.1091/mbc.E10-12-0938 (2011).
-
Ohashi, E. et al. Identification of a novel REV1-interacting motif necessary for Dna polymerase kappa function. Genes Cells xiv, 101–111. https://doi.org/x.1111/j.1365-2443.2008.01255.10 (2009).
-
Boehm, Due east. Thousand. et al. The proliferating cell nuclear antigen (PCNA)-interacting poly peptide (PIP) motif of Dna polymerase eta mediates its interaction with the C-terminal domain of Rev1. J. Biol. Chem. 291, 8735–8744. https://doi.org/10.1074/jbc.M115.697938 (2016).
-
Boehm, Eastward. M. & Washington, 1000. T. RIP to the PIP: PCNA-binding motif no longer considered specific: PIP motifs and other related sequences are non singled-out entities and can demark multiple proteins involved in genome maintenance. BioEssays 38, 1117–1122. https://doi.org/10.1002/bies.201600116 (2016).
-
Durando, Chiliad., Tateishi, Due south. & Vaziri, C. A non-catalytic function of Dna polymerase eta in recruiting Rad18 and promoting PCNA monoubiquitination at stalled replication forks. Nucleic Acids Res. 41, 3079–3093. https://doi.org/10.1093/nar/gkt016 (2013).
-
Nelson, J. R., Gibbs, P. E., Nowicka, A. Yard., Hinkle, D. C. & Lawrence, C. W. Evidence for a second function for Saccharomyces cerevisiae Rev1p. Mol. Microbiol. 37, 549–554 (2000).
-
Friedberg, Eastward. C., Lehmann, A. R. & Fuchs, R. P. Trading places: how do DNA polymerases switch during translesion DNA synthesis?. Mol. Cell xviii, 499–505. https://doi.org/10.1016/j.molcel.2005.03.032 (2005).
-
Zhou, Y., Wang, J., Zhang, Y. & Wang, Z. The catalytic function of the Rev1 dCMP transferase is required in a lesion-specific manner for translesion synthesis and base damage-induced mutagenesis. Nucleic Acids Res. 38, 5036–5046. https://doi.org/10.1093/nar/gkq225 (2010).
-
Guo, C. et al. REV1 protein interacts with PCNA: significance of the REV1 BRCT domain in vitro and in vivo. Mol. Prison cell 23, 265–271. https://doi.org/10.1016/j.molcel.2006.05.038 (2006).
-
Otsuka, C., Kunitomi, N., Iwai, Due south., Loakes, D. & Negishi, K. Roles of the polymerase and BRCT domains of Rev1 protein in translesion Deoxyribonucleic acid synthesis in yeast in vivo. Mutat. Res. 578, 79–87. https://doi.org/10.1016/j.mrfmmm.2005.03.005 (2005).
-
Guo, C. et al. Ubiquitin-binding motifs in REV1 protein are required for its role in the tolerance of DNA damage. Mol. Cell Biol. 26, 8892–8900. https://doi.org/10.1128/MCB.01118-06 (2006).
-
Guo, C. et al. Mouse Rev1 protein interacts with multiple DNA polymerases involved in translesion DNA synthesis. EMBO J. 22, 6621–6630. https://doi.org/10.1093/emboj/cdg626 (2003).
-
Ohashi, Due east. et al. Interaction of hREV1 with iii human Y-family unit Deoxyribonucleic acid polymerases. Genes Cells 9, 523–531. https://doi.org/10.1111/j.1356-9597.2004.00747.ten (2004).
-
Wojtaszek, J. et al. Multifaceted recognition of vertebrate Rev1 by translesion polymerases zeta and kappa. J. Biol. Chem. 287, 26400–26408. https://doi.org/x.1074/jbc.M112.380998 (2012).
-
Xie, W., Yang, X., Xu, M. & Jiang, T. Structural insights into the assembly of homo translesion polymerase complexes. Poly peptide Cell 3, 864–874. https://doi.org/10.1007/s13238-012-2102-x (2012).
-
Niu, Ten. et al. Rev1 plays central roles in mammalian Dna-damage tolerance in response to UV irradiation. FEBS J. 286, 2711–2725. https://doi.org/10.1111/febs.14840 (2019).
-
Qin, Z. et al. Dna-impairment tolerance mediated past PCNA*Ub fusions in human cells is dependent on Rev1 merely non Poleta. Nucleic Acids Res. 41, 7356–7369. https://doi.org/10.1093/nar/gkt542 (2013).
-
Pozhidaeva, A. et al. NMR structure and dynamics of the C-final domain from human Rev1 and its circuitous with Rev1 interacting region of Dna polymerase eta. Biochemistry 51, 5506–5520. https://doi.org/10.1021/bi300566z (2012).
-
Wojtaszek, J. et al. Structural basis of Rev1-mediated assembly of a quaternary vertebrate translesion polymerase complex consisting of Rev1, heterodimeric polymerase (Pol) zeta, and Political leader kappa. J. Biol. Chem. 287, 33836–33846. https://doi.org/10.1074/jbc.M112.394841 (2012).
-
Diamant, N. et al. DNA damage featherbed operates in the S and G2 phases of the prison cell wheel and exhibits differential mutagenicity. Nucleic Acids Res. 40, 170–180. https://doi.org/10.1093/nar/gkr596 (2012).
-
Lehmann, A. R. et al. Translesion synthesis: Y-family polymerases and the polymerase switch. Dna Repair (Amst.) 6, 891–899. https://doi.org/x.1016/j.dnarep.2007.02.003 (2007).
-
Cong, L. et al. Multiplex genome technology using CRISPR/Cas systems. Science 339, 819–823. https://doi.org/10.1126/science.1231143 (2013).
-
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 organization. Nat. Protoc. 8, 2281–2308. https://doi.org/10.1038/nprot.2013.143 (2013).
-
Yang, L., Yang, J. 50., Byrne, South., Pan, J. & Church, G. M. CRISPR/Cas9-Directed Genome Editing of Cultured Cells. Curr Protoc Mol Biol 107, 31 31 31–17, doi:https://doi.org/x.1002/0471142727.mb3101s107 (2014).
-
Moldovan, G. 50., Pfander, B. & Jentsch, S. PCNA, the maestro of the replication fork. Cell 129, 665–679. https://doi.org/x.1016/j.cell.2007.05.003 (2007).
-
Waters, L. S. et al. Eukaryotic translesion polymerases and their roles and regulation in DNA damage tolerance. Microbiol. Mol. Biol. Rev. 73, 134–154. https://doi.org/10.1128/mmbr.00034-08 (2009).
-
Ito, W. et al. Stalled Poleta at its cognate substrate initiates an culling translesion synthesis pathway via interaction with REV1. Genes Cells: Devot. Mol. Cell. Mech. 17, 98–108. https://doi.org/10.1111/j.1365-2443.2011.01576.x (2012).
-
Baldeck, N. et al. FF483-484 motif of human being Poleta mediates its interaction with the POLD2 subunit of Poldelta and contributes to DNA harm tolerance. Nucleic Acids Res. 43, 2116–2125. https://doi.org/x.1093/nar/gkv076 (2015).
-
Kim, N., Mudrak, S. V. & Jinks-Robertson, S. The dCMP transferase activity of yeast Rev1 is biologically relevant during the bypass of endogenously generated AP sites. Dna Repair (Amst.) ten, 1262–1271. https://doi.org/10.1016/j.dnarep.2011.09.017 (2011).
-
Wiltrout, M. E. & Walker, G. C. The DNA polymerase activity of Saccharomyces cerevisiae Rev1 is biologically meaning. Genetics 187, 21–35. https://doi.org/x.1534/genetics.110.124172 (2011).
-
Nair, D. T., Johnson, R. East., Prakash, L., Prakash, S. & Aggarwal, A. K. Rev1 employs a novel mechanism of DNA synthesis using a protein template. Science 309, 2219–2222. https://doi.org/10.1126/scientific discipline.1116336 (2005).
-
Zhang, Y. et al. Response of human REV1 to different DNA harm: preferential dCMP insertion contrary the lesion. Nucleic Acids Res. 30, 1630–1638. https://doi.org/10.1093/nar/30.seven.1630 (2002).
-
Temviriyanukul, P. et al. Temporally distinct translesion synthesis pathways for ultraviolet low-cal-induced photoproducts in the mammalian genome. Dna Repair (Amst.) xi, 550–558. https://doi.org/10.1016/j.dnarep.2012.03.007 (2012).
-
Gibbs, P. Due east., McDonald, J., Woodgate, R. & Lawrence, C. W. The relative roles in vivo of Saccharomyces cerevisiae Political leader eta, Politician zeta, Rev1 protein and Pol32 in the bypass and mutation induction of an abasic site, T-T (vi–4) photoadduct and T-T cis-syn cyclobutane dimer. Genetics 169, 575–582. https://doi.org/10.1534/genetics.104.034611 (2005).
-
Quinet, A. et al. Translesion synthesis mechanisms depend on the nature of DNA harm in UV-irradiated human cells. Nucleic Acids Res. 44, 5717–5731. https://doi.org/10.1093/nar/gkw280 (2016).
-
Wang, Z. & Xiao, W. Distinct requirements for budding yeast Rev1 and Poleta in translesion DNA synthesis across different types of Dna damage. Curr. Genet. 66, 1019–1028. https://doi.org/10.1007/s00294-020-01092-due west (2020).
-
Pustovalova, Y. et al. Interaction betwixt the Rev1 C-Final domain and the PolD3 subunit of Polzeta suggests a mechanism of polymerase exchange upon Rev1/Polzeta-dependent translesion synthesis. Biochemistry 55, 2043–2053. https://doi.org/10.1021/acs.biochem.5b01282 (2016).
-
Lee, Y. S., Gregory, M. T. & Yang, W. Human Pol zeta purified with accessory subunits is agile in translesion Dna synthesis and complements Pol eta in cisplatin featherbed. Proc. Natl. Acad. Sci. U.s. 111, 2954–2959. https://doi.org/10.1073/pnas.1324001111 (2014).
-
Nelson, J. R., Lawrence, C. W. & Hinkle, D. C. Deoxycytidyl transferase action of yeast REV1 poly peptide. Nature 382, 729–731. https://doi.org/x.1038/382729a0 (1996).
-
Bastien, N., Therrien, J. P. & Drouin, R. Cytosine containing dipyrimidine sites tin can exist hotspots of cyclobutane pyrimidine dimer germination after UVB exposure. Photochem. Photobiol. Sci. 12, 1544–1554. https://doi.org/10.1039/c3pp50099c (2013).
Acknowledgements
The authors wish to thank Dr. Caixia Guo for the mRev1 construct, Zhaoyan Li for technical assistance and Michelle Hanna for proofreading the manuscript. This work is supported by the National Natural Science Foundation of China operating grant 31670068 to WX.
Writer information
Affiliations
Contributions
T.B., X.Northward. and W.X. conceived and designed the experiments and wrote the manuscript. T.B., 10.N. and C.Q. performed experiments and analyzed the information. All authors read and approved the concluding manuscript.
Respective author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This commodity is licensed under a Creative Commons Attribution 4.0 International License, which permits apply, sharing, adaptation, distribution and reproduction in any medium or format, as long as y'all give appropriate credit to the original author(s) and the source, provide a link to the Creative Eatables licence, and indicate if changes were made. The images or other third party textile in this commodity are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If textile is not included in the article'south Artistic Commons licence and your intended apply is not permitted by statutory regulation or exceeds the permitted use, you will demand to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/iv.0/.
Reprints and Permissions
About this article
Cite this commodity
Bi, T., Niu, X., Qin, C. et al. Genetic and physical interactions between Polη and Rev1 in response to UV-induced Deoxyribonucleic acid impairment in mammalian cells. Sci Rep xi, 21364 (2021). https://doi.org/10.1038/s41598-021-00878-iii
-
Received:
-
Accustomed:
-
Published:
-
DOI : https://doi.org/ten.1038/s41598-021-00878-3
Comments
Past submitting a comment you concord to abide by our Terms and Community Guidelines. If you find something calumniating or that does non comply with our terms or guidelines please flag it every bit inappropriate.
Source: https://www.nature.com/articles/s41598-021-00878-3
0 Response to "Trading Places Review How Do Dna Polymerases Switch During Translesion Dna Synthesis?"
Post a Comment